Noireaux
Lab
What,
Why and How?
My
lab
has developed
a cell-free transcription-translation (TXTL) platform to construct
biochemical systems in vitro by executing synthetic gene circuits.
Unlike the other
cell-free expression systems, our platform is based on an E. coli
extract
that uses the endogenous TX and TL machineries. The circuits (plasmids
or linear DNA) are executed in a cell-free TXTL mix entirely prepared
in
our lab. This cell-free system is available under the name
myTXTL
at
Arbor Biosciences. Our
research is based on this unique system and includes: (I) prototyping
regulatory elements and
circuits, (II) quantitative biology of self-assembly with
phages as models, (III) bottom-up
construction of a minimal cell, (IV) application to
biotechnologies and medicine. Our work is both fundamental and
applied and covers the
research areas of synthetic biology and quantitative biology such as
biological physics.
Cell-free
expression systems
Cell-free
protein synthesis was developed in the 60s to understand the
process of protein synthesis in living organisms. In vitro protein
synthesis had an immediate impact with the elucidation of the genetic
code (1). In the 70s, DNA-dependent cell-free expression became
a research tool to analyze gene products and to unravel the regulation
of natural genetic elements such as the E. coli lactose (2) and
tryptophan (3) operons. The development of highly efficient hybrid
cell-free expression systems in the early 90s marked a turning point
for this technology (4). Cell-free TXTL
systems, optimized for large-scale protein synthesis as an
alternative to the recombinant protein technology (5, 6), are used in
an increasing number of applications in biotechnology, industry and
proteomics (7-9).
With
the emergence of synthetic biology, a new generation of cell-free TXTL
systems has been engineered. The construction of biological systems in
test tubes using DNA programs provides a means to study biochemical
processes in isolation, with a greater level of control and a greater
freedom of design compared to in vivo. In addition to increasing our
knowledge of the molecular repertoire found in biology, constructing
information-based biochemical systems in vitro offers the possibility
of expanding the capabilities of existing biological systems (10).
Elementary gene circuits (11, 12), pattern formation (13) and
prototypes of artificial cells (14, 15) have been engineered with
cell-free TXTL systems. Cell-free synthetic biology is a rapidly
growing
research area.

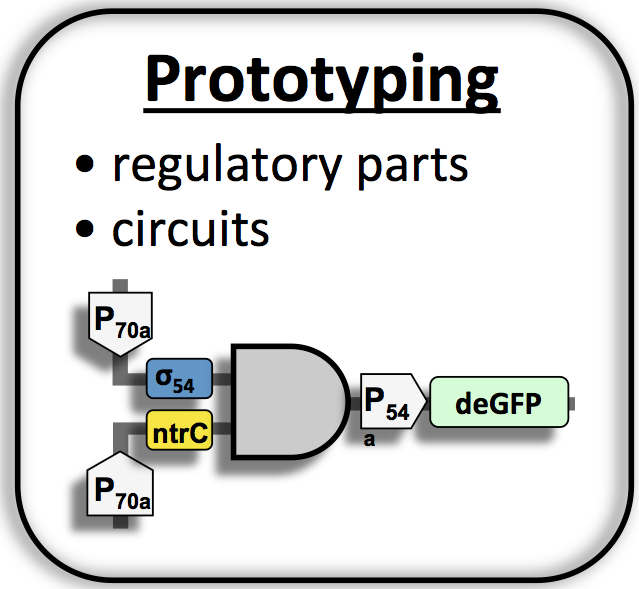
(I)
Prototyping
First,
we
demonstrated that cell-free TXTL using the E. coli endogenous
TX machinery (core RNAP + housekeeping sigma factor 70) is as
efficient as the conventional T7 bacteriophage systems (16). Methods
to tune the mRNA and protein
degradation rates were added to this system (17) so as to change the
dynamics of expression. A model of cell-free protein synthesis was
published (18), in collaboration with the Bar-Ziv lab at the
Weizmann Institute of Sciences. We then developed a
platform
that recapitulates the entire transcription scheme of E. coli (12). The
primary sigma factor 70 is used to cascade any of the
six other sigma factors 19, 24, 28, 32, 38, 54-NtrC, as well the T7 and
T3 RNA polymerases. Hundreds of
circuit parts are available from E. coli to design, build and test
synthetic circuits in vitro. We developed new metabolisms to energize
TXTL up to 2 mg/ml in batch mode (19) and published an improved TXTL
toolbox 2.0 (20). The
cell-free TXTL toolbox 2.0 is used to prototype regulatory elements
(such
as riboregulators or CRISPR guide RNAs) and gene circuits in vitro,
such as the multiple stage cascade shown in Figure 1.
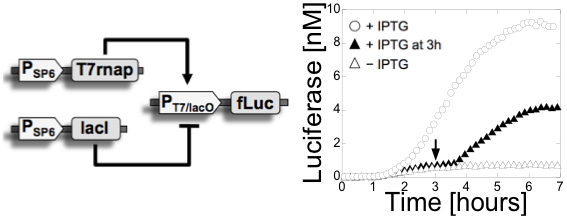
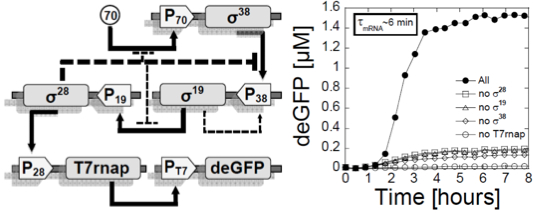
Figure
1: Left: a switch was
constructed with
bacteriophage promoters using a commercial kit. The circuit is composed
of 3 plasmids and two stages. In the first stage, the phage T7 RNA
polymerase and the E. coli lac repressor are expressed from SP6
promoters (The phage SP6 RNA polymerase is added to the TXTL
reaction). In the second stage, the expression of the luminescent
reporter protein firefly Luciferase is activated by the T7 RNA
polymerase and/or repressed by the lac repressor. The repression can be
inhibited by addition of IPTG, an inhibitor of the lac
repressor. This circuit was published in (11). Right: a multiple stage cascade
composed of 5 stages. This circuit was constructed with the new all
E. coli cell-free TXTL system specifically developed for cell-free
synthetic biology. The solid arrows show the cascades, the dotted lines
represent the negative feedback due to the competition between sigma
factors. This
circuit was published in (12) and (20).
(II)
Self-Assembly
Recently,
we challenged the system by testing large natural DNA programs. First,
we
determined that the maximum size of the genetic DNA program that can be
executed in
a test tube is about 150-250 genes, based on the toolbox 2.0
performances (20). We
expressed the T7 phage (40 kbp,
about 60 genes) and
observed its complete synthesis. In addition we showed that the phage
DNA
genome is replicated (21). It is the first time that a living entity
was entirely synthesized in vitro from the expression of its genome.
This work is not limited to the bacteriophage T7, the phages MS2 and
phiX174 can
also be synthesized in vitro (20).
.
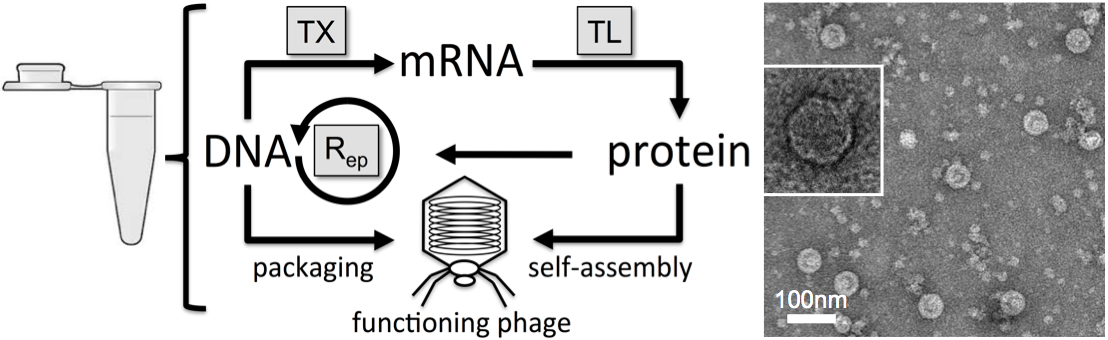
Figure
2: Left: a schematic
that
recapitulates the steps of phage expression and synthesis in a test
tube. The phage T7 has its own RNA polymerase and its own DNA
polymerase. We demonstrated that the phage T7 genome was replicated
concurrently with its synthesis. This work was published
in (20) and (21). Right:
an electron
microscopy image of T7 phages synthesized in cell-free TXTL reactions
(III)
Synthetic cell
One of the most challenging goals in cell-free synthetic biology is the
bottom up
construction minimal cells. Different types of cell analogs have
been proposed. Our approach consists of encapsulating the TXTL system
into cell-sized synthetic liposomes. The liposomes are programmed with
gene circuits towards self-reproduction, by achieving cell functions
such as
membrane permeability (14) and
cytoskeleton (22). This approach helps us understand the links
between information, self-assembly and metabolism (23).
.
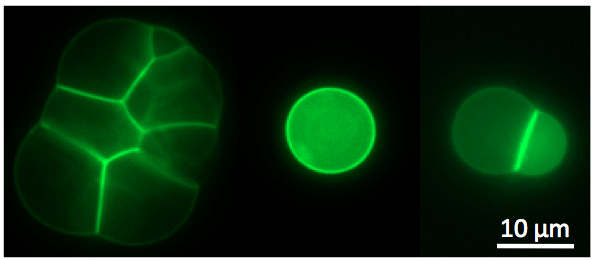
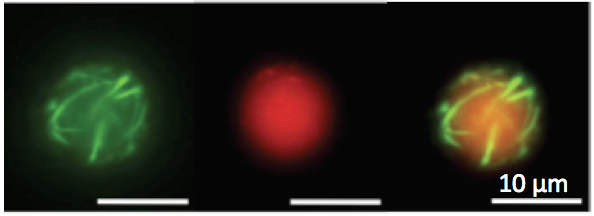
Figure
3: Left: cell-free TXTL
was used to express the toxin
alpha-hemolysin into cell-sized liposomes. The reporter protein eGFP
was fused to the toxin to visualized the interaction of the
pore-forming protein with
the phospholipid membrane. The toxin forms a membrane channel of 1.3 nm
diameter that allows exchanges of small nutrients and reaction
byproducts between the liposome and the external medium, which results
in
the extension of TXTL expression inside the liposomes. This work
was described in (14) and (20). Right:
the all E. coli toolbox was used to express the MreB and MreC
cytoskeletal protein inside liposomes at the same time. MreB
polymerizes at the membrane
through its interaction with MreC, a membrane protein. These
observations were published in (19).
(IV)
Biotechnology and Medicine
In
collaboration with other labs and companies, we are using our TXTL
system to develop molecules and devices, such as solid state probes,
for application in
biotechnologies and medicine. We use the fast prototyping capabilities
of TXTL to test and select peptides and proteins with therapeutic
functions.

1.
Nirenberg, M. (2004) Historical review:
Deciphering the
genetic
code--a personal account, Trends Biochem Sci 29, 46-54.
2. Chambers, D. A., and Zubay, G. (1969) The
stimulatory
effect of
cyclic cAMP on DNA-directed synthesis of
beta-galactosidase in a cell-free system, Proc Natl Acad Sci U S A 63,
118-122.
3. Zalkin, H., Yanofsky, C., and Squires, C. L.
(1974)
Regulated in
vitro synthesis of Escherichia coli tryptophan operon messenger
ribonucleic acid and enzymes, J Biol Chem 249, 465-475.
4. Nevin, D. E., and Pratt, J. M. (1991) A
coupled in vitro
transcription-translation system for the exclusive synthesis of
polypeptides expressed from the T7 promoter, FEBS Let 291, 259-263.
5. Kigawa, T., Yabuki, T., Yoshida, Y.,
Tsutsui, M., Ito,
Y., Shibata,
T., and Yokoyama, S. (1999) Cell-free production and stable-isotope
labeling of milligram quantities of proteins, FEBS Let 442, 15-19.
6. Jewett, M. C., and Swartz, J. R. (2004)
Rapid expression
and
purification of 100 nmol quantities of active protein using cell-free
protein synthesis, Biotechnol Prog 20, 102-109.
7. Spirin, A. S. (2004) High-throughput
cell-free systems
for synthesis
of functionally active proteins, Trends Biotechnol 22, 538-545.
8. Katzen, F., Chang, G., and Kudlicki, W.
(2005) The past,
present and
future of cell-free protein synthesis, Trends Biotechnol 23, 150-156.
9. Swartz, J. (2006) Developing cell-free
biology for
industrial
applications, J Ind Microbiol Biotechnol 33, 476-485.
10. Hodgman, C. E., and Jewett, M. C. (2012) Cell-free
synthetic
biology: Thinking outside the cell, Metab Eng, 14(3) 261-269
11. Noireaux, V., Bar-Ziv, R., and Libchaber, A. (2003)
Principles of
cell-free genetic circuit assembly, Proc Natl Acad Sci U S A 100,
12672-12677.
12. Shin, J., Noireaux, V. (2011) An E. coli cell-free
expression
toolbox: application to synthetic gene circuits and artificial cells.,
ACS Synthetic Biology 1, 29-41.
13. Isalan, M., Lemerle, C., and Serrano, L. (2005)
Engineering gene
networks to emulate Drosophila embryonic pattern formation, PLoS Biol
3(3), e64.
14. Noireaux, V., and Libchaber, A. (2004) A vesicle
bioreactor as a
step toward an artificial cell assembly, Proc Natl Acad Sci U S A 101,
17669-17674.
15. Ishikawa, K., Sato, K., Shima, Y., Urabe, I., and Yomo,
T. (2004)
Expression of a cascading genetic network within liposomes, FEBS Lett
576, 387-390.
16. Shin, J., Noireaux, V. (2010) Efficient cell-free
expression with
the endogenous E. Coli RNA polymerase and sigma factor 70, J. Biol.
Eng. 4:8.
17. Shin, J., Noireaux, V. (2010) Study of messenger RNA
inactivation
and protein degradation in an Escherichia coli cell-free expression
system, J. Biol. Eng. 4:9.
18. Karzbrun, E., Shin, J., Bar-Ziv, R., Noireaux, V. (2011)
Coarse grained dynamics of protein synthesis in a cell-free system, PRL
106(4), 048104.
19. Caschera, F. and Noireaux, V. (2013) Synthesis of 2.3 mg/ml
of protein with an all Escherichia coli cell-free
transcription-translation system. Biochimie 99, 162-168.
20. Garamella, J., Marshall, R., Rustad, M.,
Noireaux, V. (2016) The all E. coli Cell-free TX-TL Toolbox 2.0: a
platform for cell-free synthetic biology. ACS Synthetic Biology DOI:
10.1021/acssynbio.5b00296.
21. Shin, J., Jardine, P., Noireaux, V. (2012) Genome
replication,
synthesis and assembly of the bacteriophage T7 in a single cell-free
reaction, ACS Synthetic Biology 1(9), 408-413.
22. Maeda, Y., Nakadai, T., Shin, J., Uryu, K., Noireaux,
V.,
Libchaber, A.
(2012) Assembly of MreB Filaments on Vesicular Membranes: A Synthetic
Biology Approach. ACS Synthetic Biology 1(2), 53-59.
23. Noireaux, V., Maeda, Y., Libchaber, A.
(2011) Development of an artificial cell, from self-organization to
computation and self-reproduction. Proc. Nat. Acad. Sci. USA 108(9),
3473-3480.
Noireaux
lab is or was sponsored by: DARPA, ONR, NSF, BSF, HFSP and UMN.